Pre-Processing
Processing Structure
File Structure
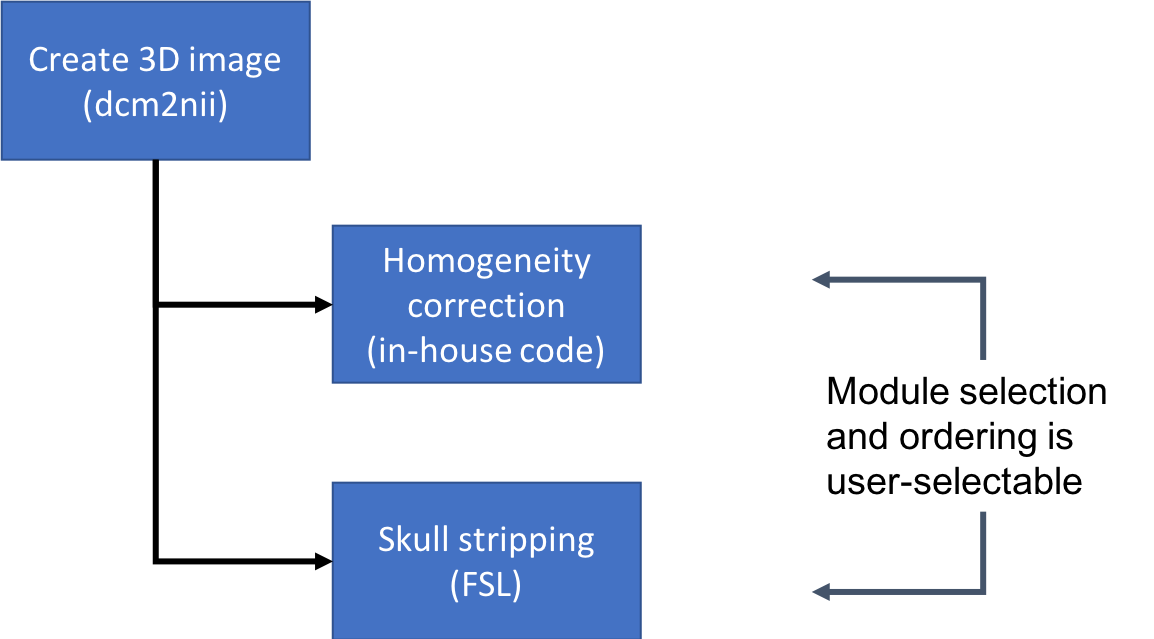
Anatomy
Subdirectories for each structural series (e.g. "t1sag_208sl")
3D structural image in "rawest" form, converted from scanner DICOM files.
Image with in-house homogeneity correction applied.
Skull-stripped image using FSL BET.
Compressed directory containing DICOM files for this series.
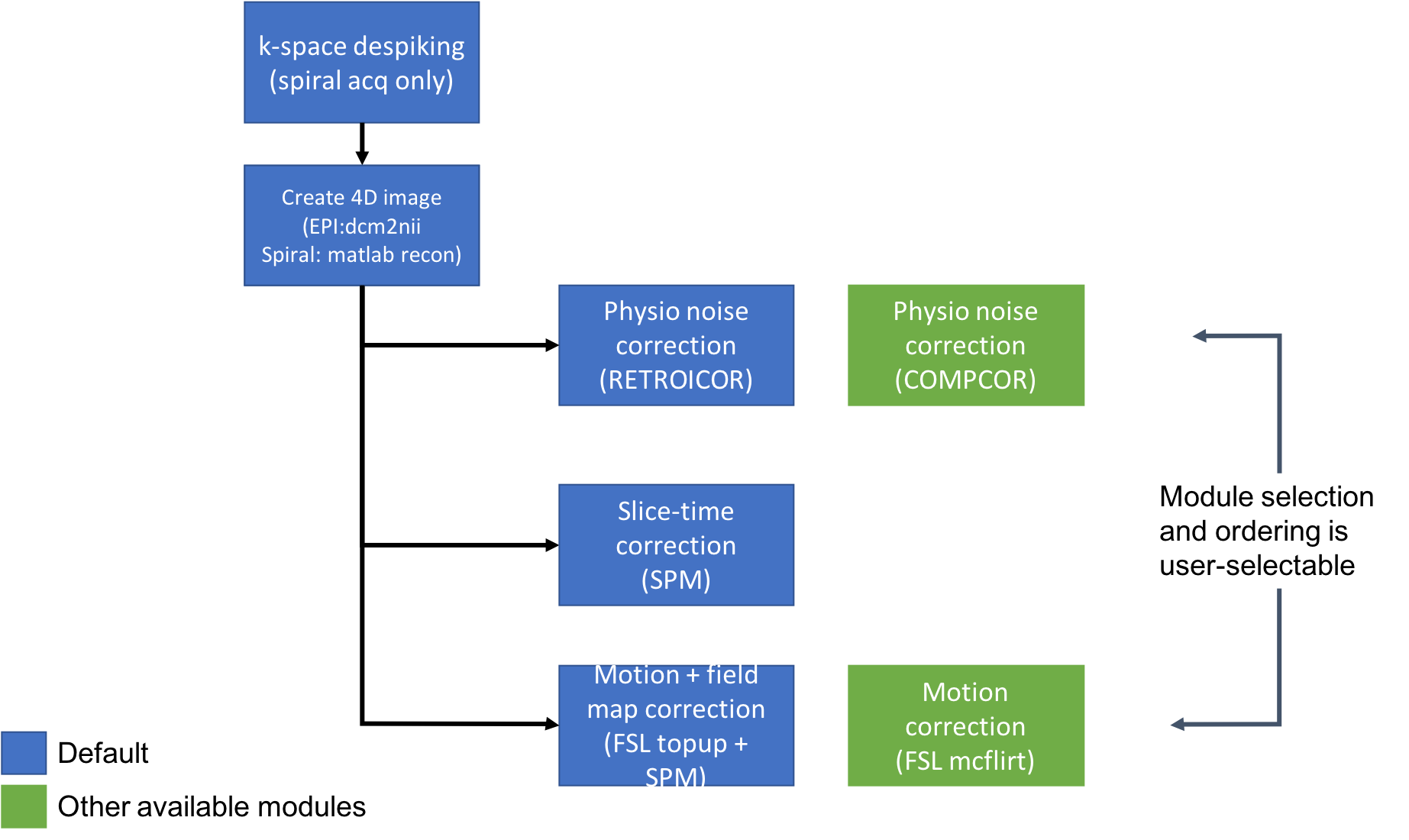
Functional (run)
Subdirectories organized as '[TASK]/[run_##]/'
4D time series image in "rawest" form, converted from scanner DICOM files or raw Pfile.
Note: For multiband data we discard the first 12 volumes of the acquisition.
Note: For multiband data we discard the first 12 volumes of the acquisition.
Physiological noise correction using RETROICOR, if physio data was obtained.
Variance maps before and after RETROICOR.
Regressor matrix from RETOROICOR
Slice-time correction (SPM).
Voxel displacement map calculated and used for field map correction.
Calculated translation (mm) and rotation (rad) movement parameters per volume, using the 10th volume of first run of the task as the reference.
Order of the 6 columns = [x_translation y_translation z_translation x_rotation y_rotation z_rotation]
Order of the 6 columns = [x_translation y_translation z_translation x_rotation y_rotation z_rotation]
Combined field map + motion correction. Field map is calculated with FSL topup and applied within SPM's realign_unwarp module.
Compressed directory containing DICOM files for this series.
Functional (field map)
Field map data found by extracting func/fieldmaps/fm_[TASK]/dicom.tgz
Field map DICOM files
Calculated field map (nifti format)
Calculated field map (Analyze format, required for SPM processing)
Calculated field map magnitude using FSL topup
Calculated field map magnitude skull-stripped with FSL bet
DTI
4D DTI file. Note: there are 6 extra B0 volumes at the start of the file. This is typically acquired with P->A phase encoding direction.
For multiband DTI we collect a short 'field map' run prior to DWI acquisition. This is a series of 8 B0 volumes, the first one being in the A->P phase encoding direction. That can be used in combination with a B0 in the DWI (collected P->A) to make the field map to use for distortion correction.